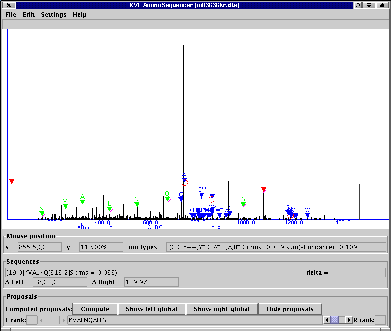
 KVL Aminosequencer permits manual interactive de novo protein
sequencing, via an easy-to-use graphical user interface.
KVL Aminosequencer permits manual interactive de novo protein
sequencing, via an easy-to-use graphical user interface.
The program reads a spectrum from a text file and displays it graphically. The program combines manual and automatic interpretation of the spectrum:
Several starting points are allowed when interpreting the spectrum. Thus, sections of the peptide can be interpreted while ambiguous parts of the spectrum are ignored. This procedure generates several short amino acid stretches per spectrum. Consequently, several database searches can easily be made of incompletely fragmented peptides. This may increase the identification percentages.
The program has been tested mainly with spectra obtained from
electrospray ionization (ESI).
Downloading and unpacking
unzip aminoseq-jar.zip
java -jar aminoseq.jar
Here are some known problems and issues:
KVL Aminosequencer was developed by Niels Jřrgen Řstergaard Kokholm (kokholm@dina.kvl.dk) and Peter Sestoft (sestoft@dina.kvl.dk).
Natascha Helena Beyer (nhbeyer@mail.dk) provided background in proteomics and comprehensive MS/MS data sets, and tested the program.
Gerhard Saalbach (g.saalbach@risoe.dk) provided the initial inspiration for this work.